DeepLagField: Time-Varying AR Reconstruction#
This tutorial demonstrates training a neural network to infer the order \(p\) and time-varying coefficients \(a(t)\) of TVAR processes:
Goals:
Generate synthetic TVAR data with known ground-truth coefficients
Train
DeepLagFieldto jointly predict AR order and coefficient trajectoriesEvaluate classification and regression accuracy across coefficient families
import torch
from torch.utils.data import DataLoader, TensorDataset
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
import numpy as np
from tqdm import tqdm
import matplotlib.pyplot as plt
import pandas as pd
from NeuralFieldManifold.models import DeepLagEmbed
from NeuralFieldManifold.generators import tvar
from NeuralFieldManifold.utils import train_loop, bench_loop
from NeuralFieldManifold.plottings import plot_history, plot_coefficients_by_p, plot_tvar_sample
from NeuralFieldManifold.generators import (
sinusoid,
fourier,
quasiperiodic,
polynomial_drift,
logistic_transition,
multi_sigmoid,
gaussian_bumps,
smooth_random,
)
from torchinfo import summary
from concurrent.futures import ProcessPoolExecutor, as_completed
import multiprocessing
from utils import pinn_processor
Coefficient Schedule Registry#
Eight families control how \(a_k(t)\) evolve over time:
sinusoid / fourier / quasiperiodic: Periodic or near-periodic oscillations
poly_drift / logistic: Smooth trends and transitions
multi_sigmoid / gaussian_bumps: Localized regime changes
smooth_random: Stochastic but differentiable trajectories
# Registry of families you can benchmark
SCHEDULES = {
"sinusoid": sinusoid,
"fourier": fourier,
"quasiperiodic": quasiperiodic,
"poly_drift": polynomial_drift,
"logistic": logistic_transition,
"multi_sigmoid": multi_sigmoid,
"gaussian_bumps": gaussian_bumps,
"smooth_random": smooth_random,
}
Data Generation#
Key functions:
simulate_tvar(): Simulates TVAR with soft power clamping to prevent divergencegenerate_one_tvar_sample(): Produces a single \((x, a(t), \text{meta})\) tuple with retry logicgenerate_dataset(): Parallelized generation balanced across families and orders \(p \in \{2,...,6\}\)
def _rng(seed):
return np.random.default_rng(seed)
def pad_to_pmax(a_time_p, p_max):
"""Pad [T, p_true] -> [T, p_max] with zeros."""
T, p = a_time_p.shape
out = np.zeros((T, p_max), dtype=a_time_p.dtype)
out[:, :p] = a_time_p
return out
def generate_one_tvar_sample(
T=10_000,
p_max=6,
schedule_name="sinusoid",
noise_std=0.3,
seed=0,
l1_cap=0.95,
P0=0.5,
W=40,
power_control=True,
p_true=None, # If provided, use this p; otherwise random
p_candidates=None,
max_retries=10000, # Maximum resampling attempts
):
rng = _rng(seed)
if p_candidates is None:
p_candidates = np.array([2, 3, 4, 5, 6], dtype=int)
# Use provided p_true or randomly select
if p_true is None:
p_true = int(rng.choice(p_candidates))
else:
p_true = int(p_true)
# Retry loop: resample until we get a valid sample
current_seed = seed
for attempt in range(max_retries):
# Use a fresh rng for coefficient schedule each attempt
attempt_rng = _rng(current_seed)
# coefficient schedule for true p
a_tp = SCHEDULES[schedule_name](T=T, p=p_true, rng=attempt_rng)
# simulate signal with power control (rejection-based)
x, a_actual = simulate_tvar(
a_tp, noise_std=noise_std, seed=current_seed+123,
P0=P0, W=W, power_control=power_control
)
# If valid sample (not rejected), break out
if x is not None:
break
# Increment seed for next attempt
current_seed += 1
else:
raise RuntimeError(f"Failed to generate valid sample after {max_retries} retries for schedule '{schedule_name}', p_true={p_true}")
# pad coeffs to p_max for consistent storage
a_t = pad_to_pmax(a_actual.astype(np.float32), p_max)
meta = {
"T": T,
"p_max": p_max,
"p_true": p_true,
"schedule": schedule_name,
"noise_std": float(noise_std),
"seed": int(current_seed), # Record the seed that actually worked
"l1_cap": float(l1_cap),
"P0": float(P0),
"W": int(W),
"power_control": power_control,
"p_candidates": list(p_candidates),
"retries": attempt, # Number of retries needed
}
return x, a_t, meta
def _generate_sample_worker(args):
"""
Worker function for parallel sample generation.
Takes a dict of arguments and returns (idx, x, a_t, meta, cid, ns).
"""
idx = args["idx"]
T = args["T"]
p_max = args["p_max"]
cname = args["cname"]
ns = args["ns"]
s = args["s"]
P0 = args["P0"]
W = args["W"]
power_control = args["power_control"]
p_true_val = args["p_true_val"]
p_candidates = args["p_candidates"]
cid = args["cid"]
x, a_t, meta = generate_one_tvar_sample(
T=T, p_max=p_max, schedule_name=cname, noise_std=ns, seed=s,
P0=P0, W=W, power_control=power_control,
p_true=p_true_val, p_candidates=p_candidates,
)
return idx, x, a_t, meta, cid, ns
def simulate_tvar(a_time, noise_std=0.3, seed=0, burn_in=300,
P0=0.5, W=40, power_control=True,
divergence_threshold=1e6):
"""
Simulate a TVAR process with soft power clamping and hard rejection safety net:
x_t = a(t)^T [x_{t-1}, ..., x_{t-p}] + eps_t
During simulation, if window power exceeds P0, the current sample is scaled
by clip(sqrt(P0/power), 0.8, 1.2) to gently steer power back toward the target.
After simulation, a final hard check rejects if power still exceeds P0.
"""
rng = _rng(seed)
T, p = a_time.shape
T_full = T + int(burn_in)
x_full = np.zeros(T_full, dtype=np.float64)
eps = rng.normal(0.0, noise_std, size=T_full)
x_full[:p] = eps[:p]
for t in range(p, T_full):
tau = t - int(burn_in)
if tau < 0:
a_t = a_time[0]
else:
a_t = a_time[min(tau, T - 1)]
lags = x_full[t-p:t][::-1]
x_full[t] = np.dot(a_t, lags) + eps[t]
# Early divergence check — fail fast instead of letting values blow up
if abs(x_full[t]) > divergence_threshold:
return None, None
# Soft power clamping (matches the working tvar() function)
if power_control and t >= W - 1:
window = x_full[t - W + 1:t + 1]
current_power = np.mean(window ** 2)
if current_power > 0:
s = np.clip(np.sqrt(P0 / current_power), 0.8, 1.2)
x_full[t] *= s
# Drop burn-in (safe cast now — divergent runs already returned None)
x = x_full[int(burn_in):].astype(np.float32)
# Hard rejection safety net: only check full W-sized windows
# (matches soft clamping which only activates at t >= W-1)
if power_control and len(x) >= W:
x2 = x ** 2
cumsum = np.concatenate([[0.0], np.cumsum(x2)])
ends = np.arange(W, len(x) + 1)
starts = ends - W
power = (cumsum[ends] - cumsum[starts]) / W
if np.any(power > P0):
return None, None
return x, a_time
def generate_dataset(
n_per_class=10,
classes=None,
T=1000,
noise_range=(0.1, 0.6),
seed=0,
dtype=np.float32,
P0=0.5,
W=40,
power_control=True,
p_candidates=None,
balanced_p=True, # If True, balance samples across p_candidates
n_workers=None, # Number of parallel workers (None = CPU count)
):
p_max = 10
if p_candidates is None:
p_candidates = np.array([2, 3, 4, 5, 6], dtype=int)
else:
p_candidates = np.array(p_candidates, dtype=int)
if classes is None:
classes = list(SCHEDULES.keys())
if n_workers is None:
n_workers = multiprocessing.cpu_count()
rng = _rng(seed)
N = n_per_class * len(classes)
X = np.zeros((N, T), dtype=dtype)
A = np.zeros((N, T, p_max), dtype=dtype)
p_true_arr = np.zeros((N,), dtype=np.int32)
class_id = np.zeros((N,), dtype=np.int32)
noise_std = np.zeros((N,), dtype=dtype)
class_names = list(classes)
# Precompute p assignments if balanced
n_p = len(p_candidates)
if balanced_p:
n_per_p = n_per_class // n_p
remainder = n_per_class % n_p
# Create balanced p assignments for one class
p_assignments = []
for i, p in enumerate(p_candidates):
count = n_per_p + (1 if i < remainder else 0)
p_assignments.extend([p] * count)
# Build all task arguments
tasks = []
idx = 0
for cid, cname in enumerate(class_names):
# Shuffle p assignments for this class
if balanced_p:
class_p_assignments = rng.permutation(p_assignments)
for i in range(n_per_class):
s = int(rng.integers(0, 2**31 - 1))
ns = float(rng.uniform(noise_range[0], noise_range[1]))
# Determine p_true
if balanced_p:
p_true_val = int(class_p_assignments[i])
else:
p_true_val = None # Let generate_one_tvar_sample pick randomly
tasks.append({
"idx": idx,
"T": T,
"p_max": p_max,
"cname": cname,
"ns": ns,
"s": s,
"P0": P0,
"W": W,
"power_control": power_control,
"p_true_val": p_true_val,
"p_candidates": list(p_candidates),
"cid": cid,
})
idx += 1
metas = [None] * N
# Parallel execution
with ProcessPoolExecutor(max_workers=n_workers) as executor:
futures = {executor.submit(_generate_sample_worker, task): task["idx"] for task in tasks}
with tqdm(total=N, desc=f"Generating samples ({n_workers} workers)") as pbar:
for future in as_completed(futures):
idx, x, a_t, meta, cid, ns = future.result()
X[idx] = x.astype(dtype)
A[idx] = a_t.astype(dtype)
p_true_arr[idx] = meta["p_true"]
class_id[idx] = cid
noise_std[idx] = ns
metas[idx] = meta
pbar.update(1)
dataset = {
"X": X,
"A": A,
"p_true": p_true_arr,
"class_id": class_id,
"class_names": np.array(class_names),
"noise_std": noise_std,
"meta": metas, # Python list of dicts; fine for notebook use
}
return dataset
def save_dataset_npz(path, dataset):
np.savez(
path,
X=dataset["X"],
A=dataset["A"],
p_true=dataset["p_true"],
class_id=dataset["class_id"],
class_names=dataset["class_names"],
noise_std=dataset["noise_std"],
meta=np.array(dataset["meta"], dtype=object),
)
def load_dataset_npz(path):
d = np.load(path, allow_pickle=True)
return {
"X": d["X"],
"A": d["A"],
"p_true": d["p_true"],
"class_id": d["class_id"],
"class_names": d["class_names"],
"noise_std": d["noise_std"],
"meta": list(d["meta"]),
}
Visualize Schedule Families#
Each schedule produces qualitatively different signals. Top row shows \(x(t)\); bottom row shows the true \(a_k(t)\) coefficients. Note how coefficient dynamics translate into signal structure.
Generate Training Dataset#
Create a balanced dataset: equal samples per family, uniform distribution over \(p \in \{2,3,4,5,6\}\), and noise \(\sigma \in [0.2, 0.5]\). Power control ensures bounded variance.
classes = list(SCHEDULES.keys())
# small pilot: 10 per family => 80 samples total
pilot = generate_dataset(
n_per_class=10,
classes=classes,
T=600,
noise_range=(0.2, 0.5),
seed=123,
dtype=np.float32
)
# save_dataset_npz("train_data.npz", pilot)
X_train, coef_train, p_train, class_id_train, X_val, coef_val, p_val, class_id_val, class_names = pinn_processor("train_data.npz", family=None)
Total samples: 1000000 | Train: 800000 | Val: 200000
Train p distribution: {np.int64(0): np.int64(160000), np.int64(1): np.int64(160000), np.int64(2): np.int64(160000), np.int64(3): np.int64(160000), np.int64(4): np.int64(160000)}
Val p distribution: {np.int64(0): np.int64(40000), np.int64(1): np.int64(40000), np.int64(2): np.int64(40000), np.int64(3): np.int64(40000), np.int64(4): np.int64(40000)}
Prepare DataLoaders#
Move tensors to GPU upfront to eliminate per-batch transfer overhead. The model expects shape (batch, T) for signals and (batch,) for order labels.
device = torch.device('cuda:1' if torch.cuda.is_available() else 'cpu')
if device.type == 'cuda':
torch.cuda.init()
torch.backends.cudnn.benchmark = True
print(f"Using device: {device}")
# Move ALL data to GPU upfront - eliminates CPU->GPU transfer bottleneck
X_train = torch.tensor(X_train, dtype=torch.float32, device=device)
p_train = torch.tensor(p_train, dtype=torch.long, device=device)
X_val = torch.tensor(X_val, dtype=torch.float32, device=device)
p_val = torch.tensor(p_val, dtype=torch.long, device=device)
class_id_val = class_id_val # numpy array
# Data already on GPU - no pin_memory/workers needed
train_loader = DataLoader(
TensorDataset(X_train, p_train),
batch_size=256,
shuffle=True
)
val_loader = DataLoader(
TensorDataset(X_val, p_val),
batch_size=256
)
Using device: cuda:1
Define Training Configuration#
Loss function combines:
lambda_p: Cross-entropy for order classificationlambda_ar: MSE between predicted and true coefficientslambda_energy: Penalizes large coefficient magnitudes within the prediction windowlambda_smooth: Encourages temporally smooth \(\hat{a}(t)\)
def do_bench_on_config(lambda_config, model=None):
# hyperparameters
n_epochs = 100
lr = 1e-3
max_ar_order = 6
if model is None:
# Model: n_classes=5 for p∈{2,3,4,5,6}, max_ar_order=6 for coefficient dimensions
model = DeepLagEmbed(seq_len=600, n_classes=5, max_ar_order=6, hidden_dim=128)
model = model.to(device)
# summary(model, input_size=(32, 600), device=device)
# Train
history = train_loop(
model, train_loader, val_loader,
n_epochs=n_epochs, lr=lr,
lambda_p=lambda_config["lambda_p"], lambda_ar=lambda_config["lambda_ar"], lambda_energy=lambda_config["lambda_energy"],
lambda_smooth=lambda_config["lambda_smooth"], lambda_order=lambda_config["lambda_order"],
p_max=max_ar_order, device=device
)
return model, history
Train Model#
Train for 100 epochs with balanced loss weights. The training curve shows joint convergence of classification and regression objectives.
conf = {
"lambda_ar": 5,
"lambda_p": 5,
"lambda_order": 0,
"lambda_energy": 0.2,
"lambda_smooth": 5,
}
model, history = do_bench_on_config(conf)
plot_history(history, model=model, val_loader=val_loader, device=device)
Training: 100%|██████████| 100/100 [14:56<00:00, 8.96s/it, p_acc=0.340, train=6.5934, val=8.4380]
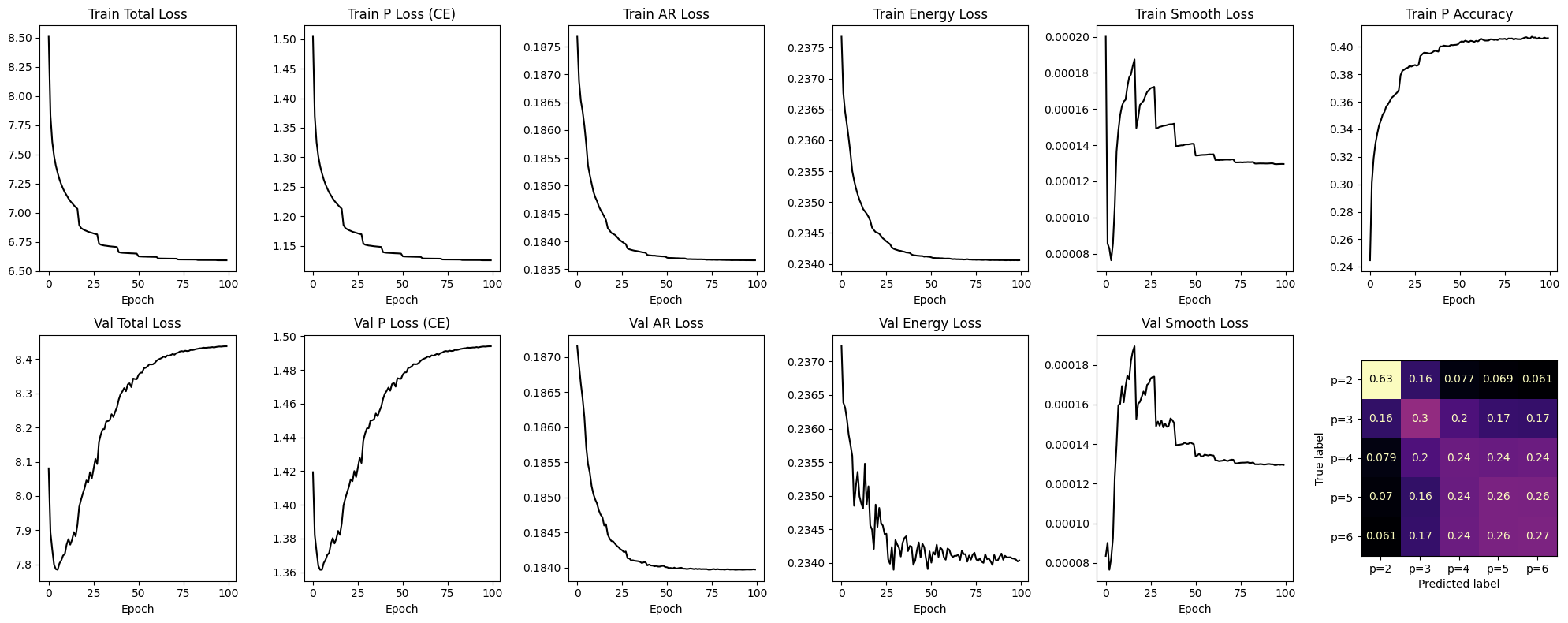
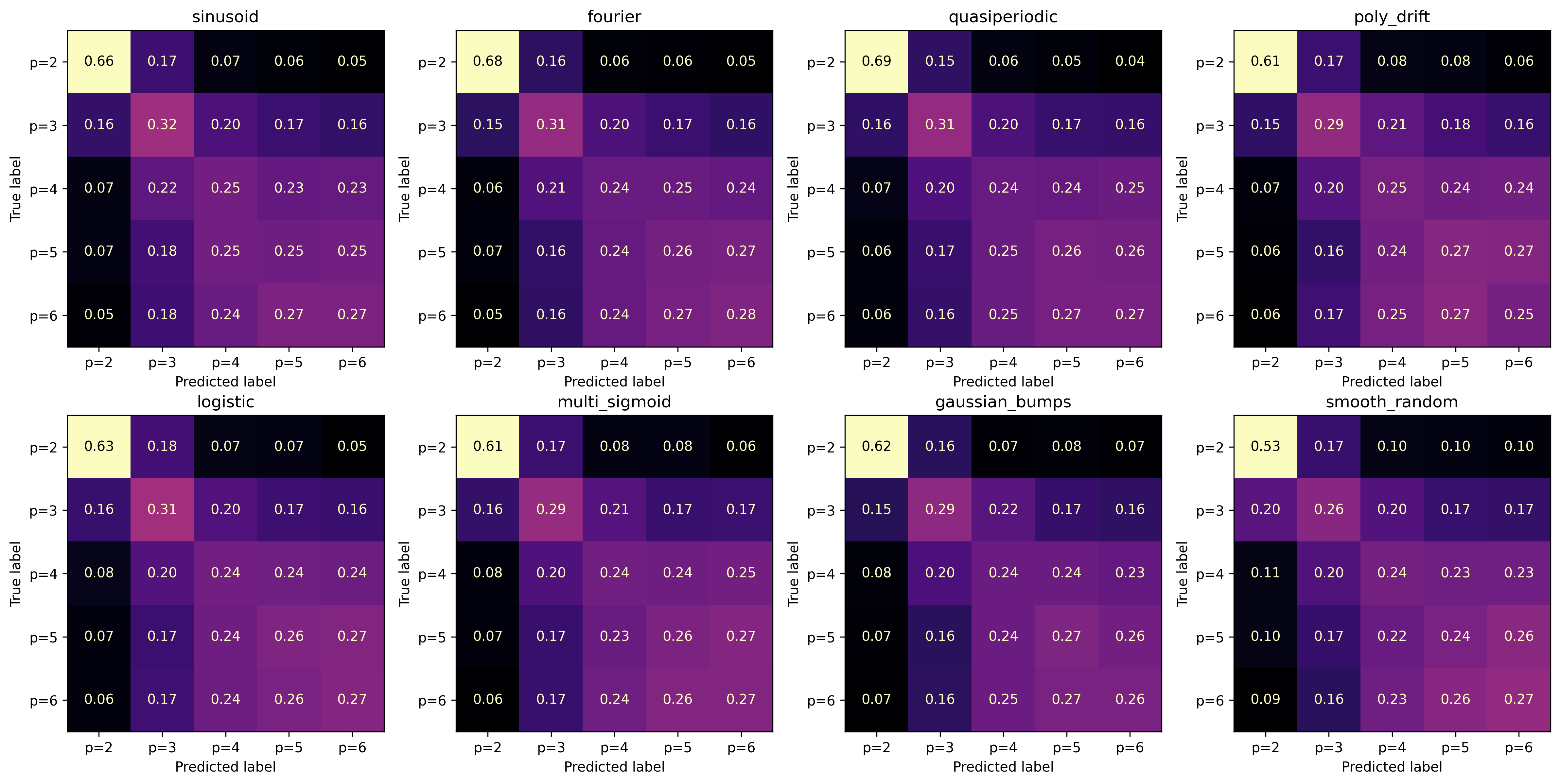
Evaluation: Order Classification#
Confusion matrices show how well the model distinguishes AR orders within each family. Diagonal dominance indicates correct classification; off-diagonal mass reveals systematic over/under-estimation of \(p\).
# Get predictions on full validation set
model.eval()
all_p_pred = []
all_p_true = []
with torch.no_grad():
for x_batch, p_batch in tqdm(val_loader, desc="Getting predictions"):
x_batch = x_batch.to(device)
_, _, p_hard, _ = model(x_batch)
all_p_pred.extend(p_hard.cpu().numpy())
all_p_true.extend(p_batch.cpu().numpy())
all_p_pred = np.array(all_p_pred)
all_p_true = np.array(all_p_true)
Getting predictions: 100%|██████████| 782/782 [00:00<00:00, 795.68it/s]
# Plot confusion matrix for each signal family
n_families = len(class_names)
nrows, ncols = 2, 4
fig, axes = plt.subplots(nrows, ncols, figsize=(4 * ncols, 4 * nrows), dpi=300)
axes_flat = axes.flatten()
p_min, p_max = 2, 6
n_classes = p_max - p_min + 1
family_results = {}
for i, family_name in enumerate(class_names):
mask = class_id_val == i
p_true_family = all_p_true[mask]
p_pred_family = all_p_pred[mask]
# Compute normalized confusion matrix
cm = confusion_matrix(p_true_family, p_pred_family, labels=range(n_classes))
cm_norm = cm / cm.sum(axis=1, keepdims=True)
# Store accuracy for this family
family_results[family_name] = {
'accuracy': np.mean(p_true_family == p_pred_family),
'n_samples': len(p_true_family)
}
# Plot
disp = ConfusionMatrixDisplay(cm_norm, display_labels=[f'p={j+p_min}' for j in range(n_classes)])
disp.plot(ax=axes_flat[i], cmap='magma', colorbar=False, values_format='.2f')
axes_flat[i].set_title(f'{family_name}')
# Hide unused subplots
for j in range(n_families, nrows * ncols):
axes_flat[j].set_visible(False)
plt.tight_layout()
plt.show()
Benchmark: Coefficient Recovery#
Metrics per family:
coeff_mse: Mean squared error of \(\hat{a}(t)\) vs true \(a(t)\)
signal_mse: One-step prediction error using estimated coefficients
p_acc: Classification accuracy for each order
family_bench_results = {}
for i, family_name in enumerate(class_names):
mask = class_id_val == i
X_family = X_val[mask].cpu() # Move to CPU for bench_loop
coef_family = coef_val[mask][:, :, :6] # Slice to max_ar_order=6
p_family = p_val[mask].cpu()
results = bench_loop(model, X_family, coef_family, p_family, device)
family_bench_results[family_name] = results
# Display as DataFrame
family_bench_df = pd.DataFrame(family_bench_results).T
family_df = family_bench_df.rename(columns={
'coeff_mse': 'coeff mse',
'signal_mse': 'signal mse',
'p_mae': 'delta p',
'p_mape': 'delta p / p',
'p2_acc': 'p=2',
'p3_acc': 'p=3',
'p4_acc': 'p=4',
'p5_acc': 'p=5',
'p6_acc': 'p=6',
'p_acc': 'P Avg.',
})
family_bench_df
| coeff_mse | signal_mse | p_mae | p_mape | p2_acc | p3_acc | p4_acc | p5_acc | p6_acc | p_acc | |
|---|---|---|---|---|---|---|---|---|---|---|
| sinusoid | 0.003924 | 0.183046 | 1.085483 | 0.290002 | 0.661209 | 0.326721 | 0.241122 | 0.251023 | 0.267625 | 0.351111 |
| fourier | 0.004272 | 0.184083 | 1.069846 | 0.284058 | 0.679984 | 0.316788 | 0.244737 | 0.274840 | 0.280372 | 0.358101 |
| quasiperiodic | 0.004035 | 0.182153 | 1.063116 | 0.281938 | 0.684039 | 0.310689 | 0.247123 | 0.272243 | 0.277012 | 0.356821 |
| poly_drift | 0.004315 | 0.183816 | 1.109580 | 0.302322 | 0.618354 | 0.300617 | 0.248848 | 0.262569 | 0.266481 | 0.340402 |
| logistic | 0.003673 | 0.183003 | 1.106807 | 0.300061 | 0.629938 | 0.296878 | 0.240559 | 0.260685 | 0.267918 | 0.339890 |
| multi_sigmoid | 0.003155 | 0.182505 | 1.132916 | 0.309419 | 0.607890 | 0.296363 | 0.244748 | 0.260765 | 0.269086 | 0.336374 |
| gaussian_bumps | 0.003542 | 0.183888 | 1.123097 | 0.302140 | 0.624358 | 0.305849 | 0.248492 | 0.266004 | 0.264642 | 0.340380 |
| smooth_random | 0.007759 | 0.189260 | 1.221906 | 0.343524 | 0.523409 | 0.258195 | 0.228875 | 0.251690 | 0.265594 | 0.305647 |
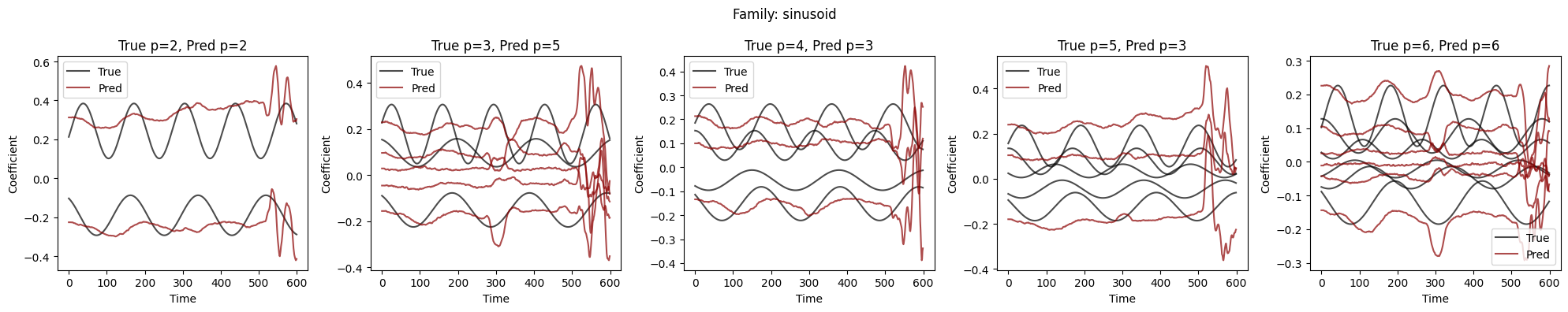
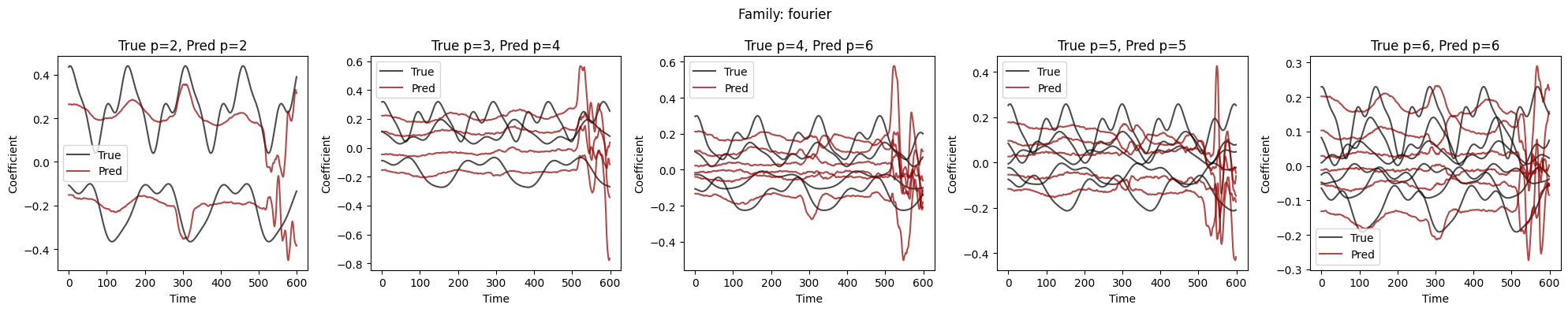
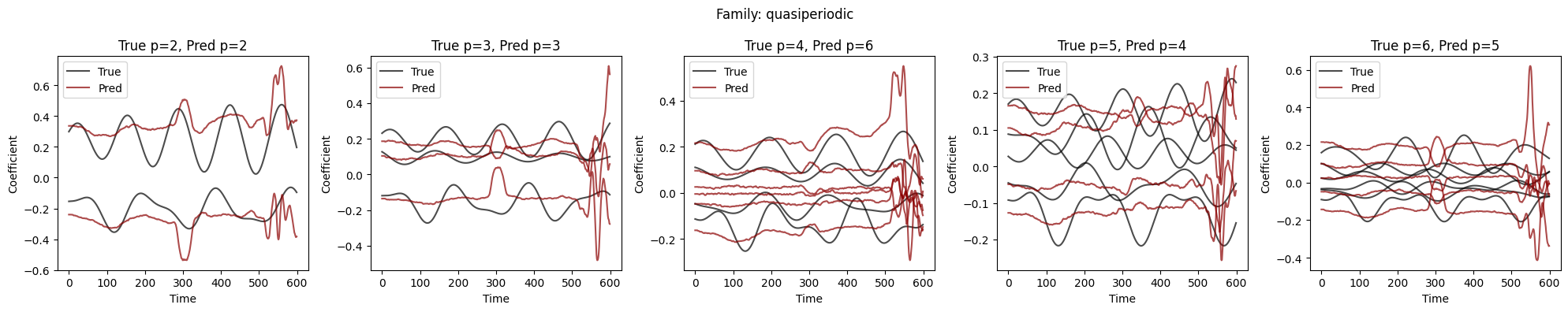
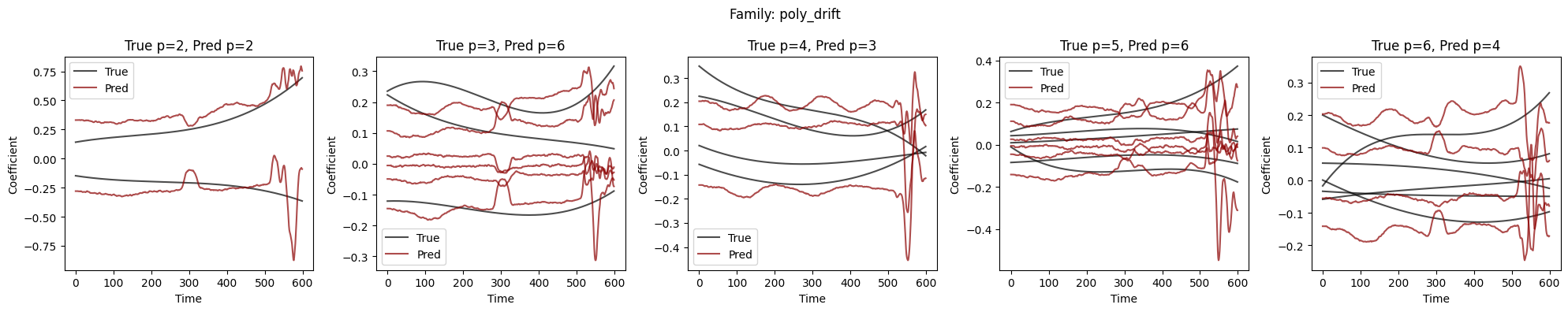
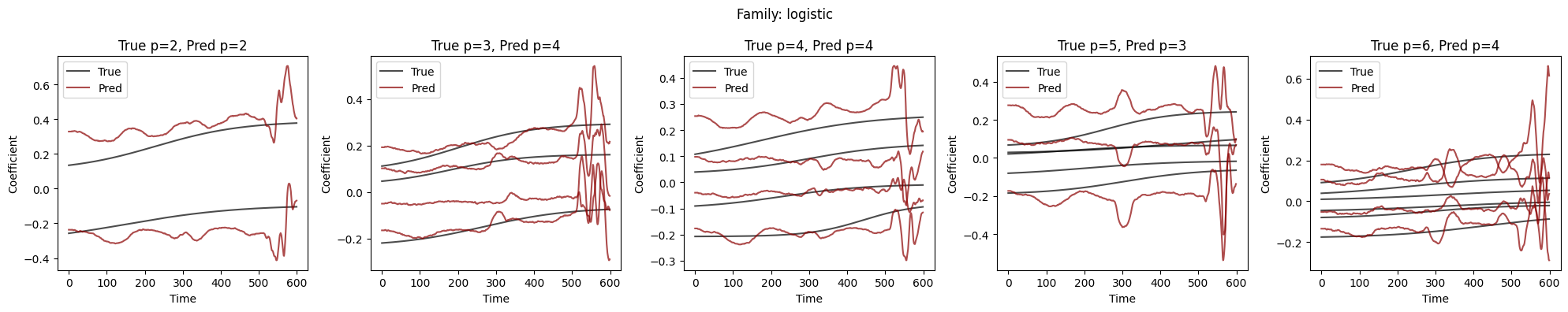
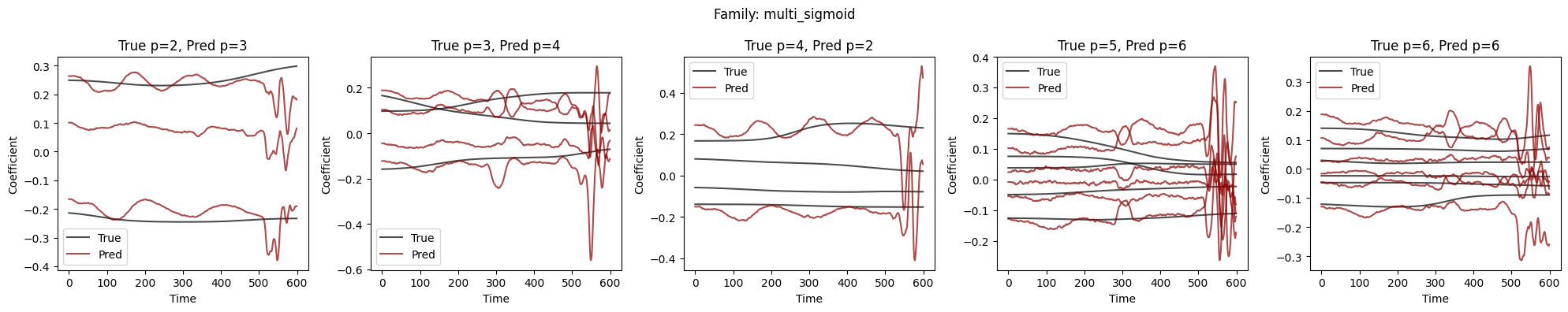
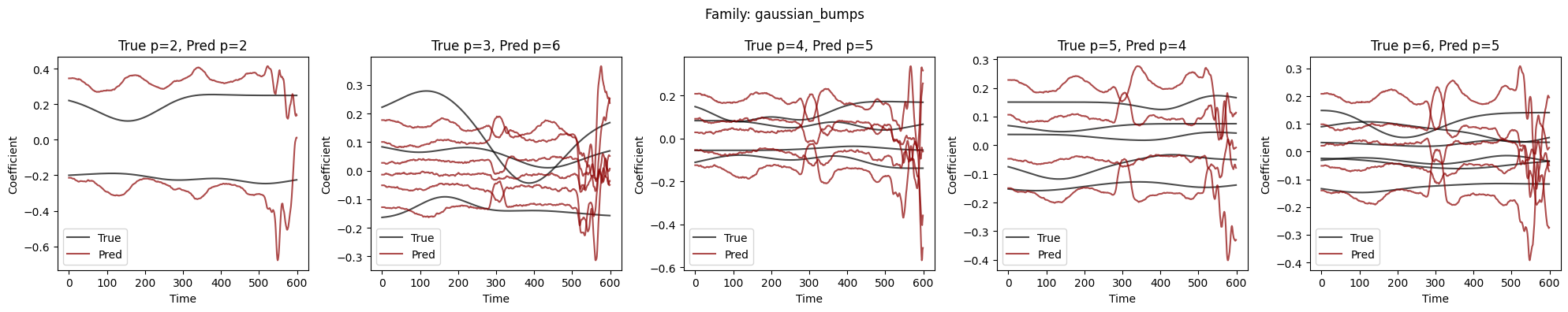
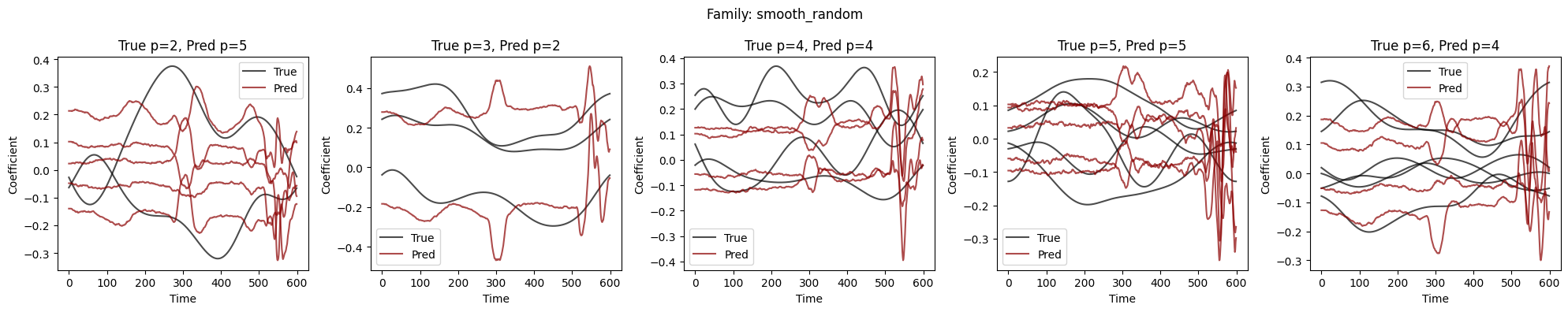
Visualize Coefficient Predictions#
Overlay of predicted \(\hat{a}_k(t)\) (dashed) and true \(a_k(t)\) (solid) for representative samples. Good fits track the ground-truth dynamics; discrepancies highlight challenging regimes.
# Plot coefficient predictions for each signal family
for i, family_name in enumerate(class_names):
mask = class_id_val == i
X_family = X_val[mask].cpu()
coef_family = coef_val[mask][:, :, :6]
p_family = p_val[mask].cpu()
plot_coefficients_by_p(model, X_family, coef_family, p_family, device,
p_max=6, p_min=2, title=f"Family: {family_name}")