Intrinsic Manifold Reconstruction and its Geometry#
import numpy as np
import mne
from scipy.ndimage import gaussian_filter1d
import matplotlib.pyplot as plt
from matplotlib.ticker import ScalarFormatter
from statsmodels.tsa.stattools import acf
from NeuralFieldManifold.generators import lorenz
from NeuralFieldManifold.models import AR, TAR
from NeuralFieldManifold.embedders import embed
from NeuralFieldManifold.pub_utils import *
set_pub_style()
Lorenz System & Spectral Content#
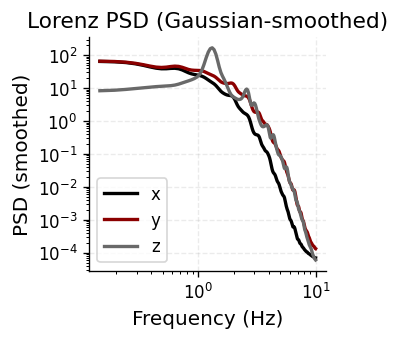
Simulate the Lorenz attractor (\(\sigma=10, \rho=28, \beta=8/3\)) and inspect its power spectral density. The broadband PSD with no sharp spectral peaks confirms chaotic, aperiodic dynamics — a useful test case for autoregressive modeling.
xs, ys, zs = lorenz(T_steps=40000,
dt=0.005,
sigma=10.0,
rho=28.0,
beta=8/3,
x0=(1.0, 1.0, 1.0))
fs = 1.0 / 0.005
burn = 2000 # drop transient
data_psd = np.vstack([xs[burn:], ys[burn:], zs[burn:]]) #(3, n_times)
# compute PSD for all three channels
psds, freqs = mne.time_frequency.psd_array_welch(
data_psd,
sfreq=fs,
fmin=0.1, fmax=10,
n_fft=4096,
n_overlap=2048,
average='mean'
)
# smooth each PSD along frequency axis
Pxx_smooth = gaussian_filter1d(psds[0], sigma=2)
Pyy_smooth = gaussian_filter1d(psds[1], sigma=2)
Pzz_smooth = gaussian_filter1d(psds[2], sigma=2)
Effective window size : 20.480 (s)
AR(p) Order Selection#
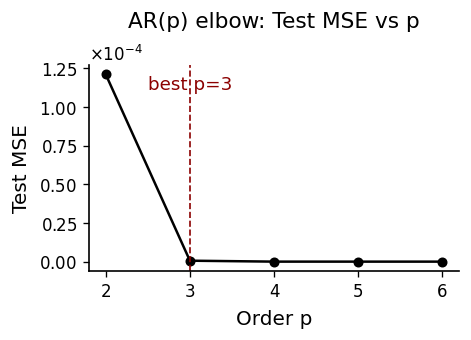
Fit univariate AR models on the \(x\)-component for orders \(p \in \{2,\dots,6\}\). We select \(p\) via test-set MSE and BIC, using an 80/20 train/test split with OLS estimation.
p_list = list(range(2, 7))
mse_te_list, aic_list, bic_list = [], [], []
params_by_p = {}
splits = {}
for p in p_list:
X, y = AR.lag_matrix(xs, p)
N = len(y)
ntr = int(0.8 * N)
X_tr, y_tr = X[:ntr], y[:ntr]
X_te, y_te = X[ntr:], y[ntr:]
w = AR.ols_with_intercept(X_tr, y_tr)
yhat_tr = AR.predict_from_params(X_tr, w)
yhat_te = AR.predict_from_params(X_te, w)
mse_te, _, _ = AR.metrics(y_te, yhat_te)
aic, bic = AR.aic_bic(y_tr, yhat_tr, k=p+1)
mse_te_list.append(mse_te); aic_list.append(aic); bic_list.append(bic)
params_by_p[p] = w
splits[p] = (ntr, y, yhat_tr, yhat_te)
best_p_by_mse = p_list[int(np.argmin(mse_te_list))]
best_p_by_bic = p_list[int(np.argmin(bic_list))]
best_p_by_mse = 3
best_p_by_bic = 3
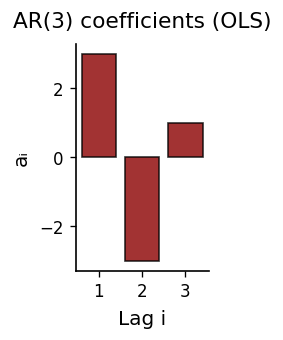
Fitted Coefficients & One-Step Prediction#
The AR(\(p\)) coefficient magnitudes show how much each lag contributes. We then evaluate the model’s one-step-ahead forecast against held-out data.
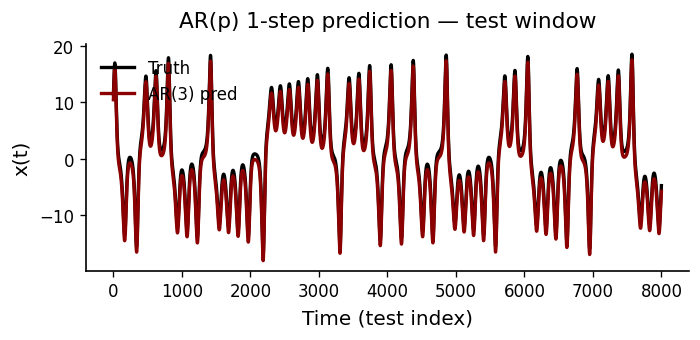
Residual Diagnostics#
If the AR model captures all linear dependence, residuals should be white noise. The ACF of test residuals should fall within the 95% confidence bands (\(\pm 1.96/\sqrt{N}\)). Significant autocorrelation at low lags signals model misspecification.
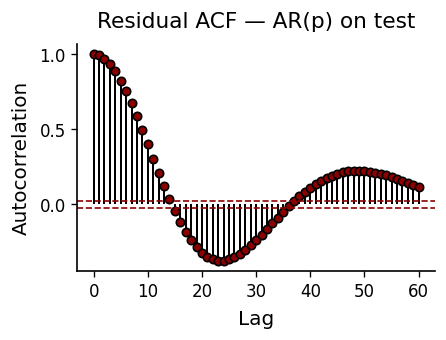
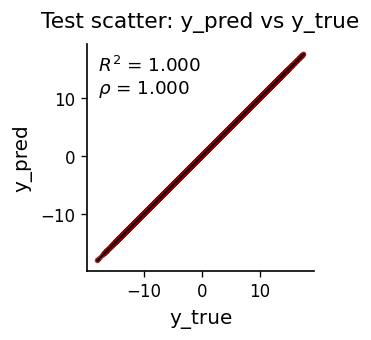
Threshold AR & Attractor Reconstruction#
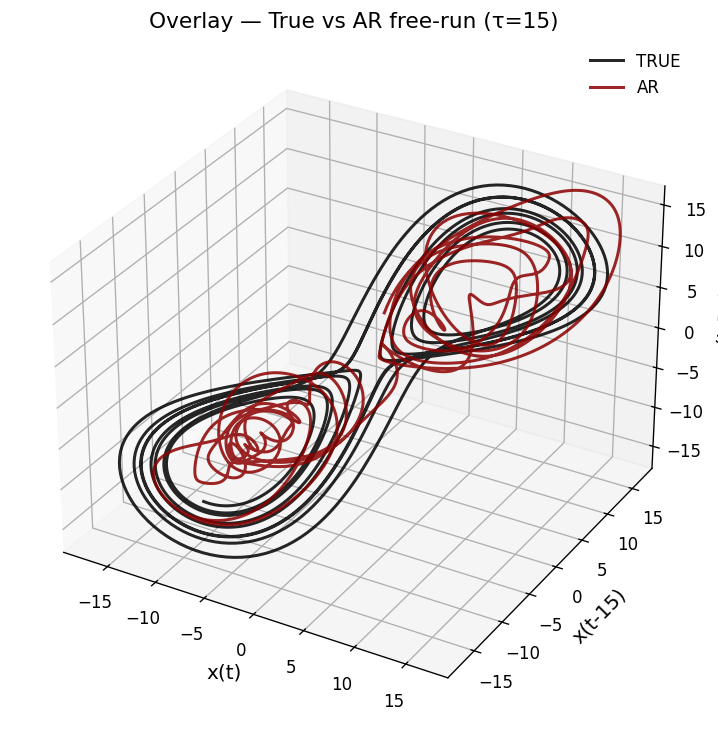
A linear AR model captures one-step dynamics but cannot reproduce the Lorenz attractor’s geometry under free-run simulation. We fit a Threshold AR (TAR) model, which switches regimes based on a delay variable, then free-run simulate and apply time-delay embedding (\(m=3, \tau=15\)) to compare the reconstructed attractors.
xs, ys, zs = lorenz(T_steps=12000)
# Fit TAR
tar = TAR.fit(xs, p=8, d=1, thresh=0.0, train_frac=0.8)
p_use, d_use = tar["p"], tar["d"]
ntr = tar["ntr"]
true_test = tar["y_all"][ntr:]
t_x_start_test = p_use + ntr
init = [xs[t_x_start_test - i] for i in range(1, p_use+1)]
n_steps = len(true_test)
# Simulate TAR with bootstrap residuals and variance matching to the test slice
x_tar = TAR.simulate_tar_free_run(tar,
init_lags=init,
n_steps=n_steps,
seed=42,
variance_match_to=true_test)
# Embeddings with τ=15
tau = 15
X3_true = embed(true_test, m=3, tau=tau)
X3_tar = embed(x_tar, m=3, tau=tau)